Research
Generative AI & Molecular Design
AI/MLDrug Discovery
A drug candidate must be potent, selective, safe, metabolically stable, and possible to synthesize. Optimizing all of these properties at once is the central challenge of molecular design. I develop generative models that tackle this multi-parameter optimization through R-group optimization, core-hopping, and knowledge-based molecular design, working directly with medicinal chemistry teams to accelerate design-make-test-analyze cycles. Early SMILES-based RNN approaches generated chemically valid but often practically inaccessible molecules. I transitioned to GFlowNet-based architectures that generate molecules exclusively over validated reaction templates using commercially available building blocks, ensuring every proposed compound carries a reviewable synthetic route. Predictive property models serve as reward functions that teach the generative models which regions of chemical space to explore, and multi-stage curriculum learning guides training from broad chemical validity toward the full multi-parameter objective.
Molecular Property Prediction
AI/MLCheminformatics
I built an automated benchmarking and deployment pipeline for predicting absorption, distribution, metabolism, excretion, and toxicity (ADMET) endpoints across multiple therapeutic programs. The pipeline spans gradient-boosted trees, message-passing neural networks, and foundation models. In my experience, careful data curation and splitting drive model quality more than architecture selection. I use clustering-based scaffold splits and temporal splits to approximate the out-of-distribution challenges of real drug discovery, and statistical hypothesis testing to provide grounded architecture selection for each endpoint. These models are deployed directly to medicinal chemists for real-time property predictions and serve as reward functions for generative molecular design. As independent external validation, I applied similar pipelines to the OpenADMET-ExpansionRx Blind Challenge and ranked 25th among 100 finalists across 9 ADMET endpoints.
Leaderboard: OpenADMET-ExpansionRx Challenge · Code: OpenADMET-ExpansionRx-Blind-Challenge
Polyelectrolyte Complexation & Ion Binding
PhysicsSimulation
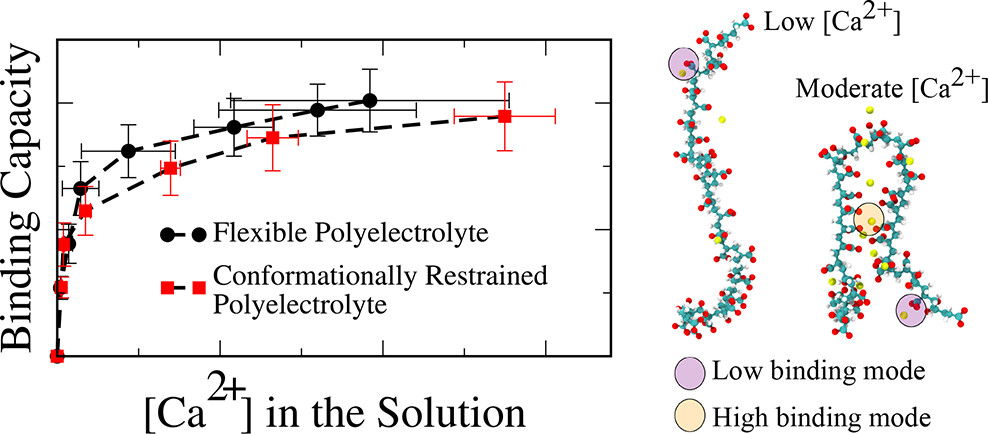
Charged polymers (polyelectrolytes) in water bind multivalent ions like Ca²⁺, a property exploited in water softening, drug delivery, and scale inhibition. Yet the molecular mechanisms underlying ion-mediated chain-chain attraction remained elusive. Using well-tempered metadynamics and Hamiltonian replica exchange, I calculated binding isotherms and free-energy landscapes for Ca²⁺ on poly(acrylic acid). Increasing Ca²⁺ concentrations induced attraction between chains, but the binding energy was not contingent on the number of ion bridges formed; instead, correlations between chelated ions were the primary driver. At high ionic strengths, electrostatic screening significantly reduced ion bridging. Surface adsorption studies showed that water-mediated hydrogen bonds and ion bridges through interfacial water, not direct polymer-surface contacts, govern binding on CaCO₃. This toolkit was extended to copolymers of acrylic acid with vinyl acetate and vinyl alcohol, and to polypeptides of aspartate and glutamate. This work was conducted in collaboration with Dow Chemical.
Key papers: Langmuir 2025 · Macromolecules 2024 · Langmuir 2024
AI/ML Methods in Soft Matter
AI/MLDeep Learning
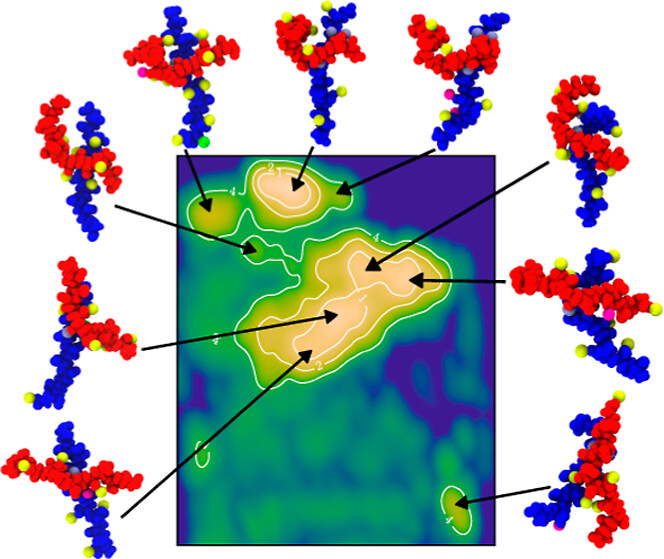
Polyelectrolyte simulations produce high-dimensional free-energy landscapes where interpretable structure-property relationships are hard to extract directly. I trained an autoencoder neural network to analyze ion-polyelectrolyte complex structures; the learned latent space effectively differentiated dominant polymer conformations and distinguished ions bridging across chains from those adsorbed on single chains, revealing distinct contributions of each binding mode. I also developed denoising diffusion probabilistic models (DDPMs) to generate polymer conformational ensembles consistent with Boltzmann statistics, accelerating conformational exploration that would otherwise require substantially more simulation time.
Code: DDPM-Enhanced-Sampling
Microhydrodynamics & Active Matter
PhysicsFluid Dynamics
Schools of fish, flocks of birds, and bacterial colonies all display spontaneous collective ordering. I set out to understand how much of this behavior emerges from fluid mechanics alone, without phenomenological interaction rules. Using a multipole expansion technique, I derived an analytical framework for self-propulsion in potential flow showing that a deformable body achieves net displacement by exploiting its configuration-dependent added mass, without performing net work on the fluid. Self-propulsion had previously been thought to require viscous dissipation, which is absent in potential flow. I then developed C++/CUDA many-body simulations parallelized via OpenMP, demonstrating that purely hydrodynamic interactions are sufficient to produce emergent collective structures. The fluid medium establishes a natural length scale, dependent on body configuration and velocity, that governs the system dynamics.
Key paper: J. Fluid Mechanics 2022

Lipid Membrane Mechanics
PhysicsTheory
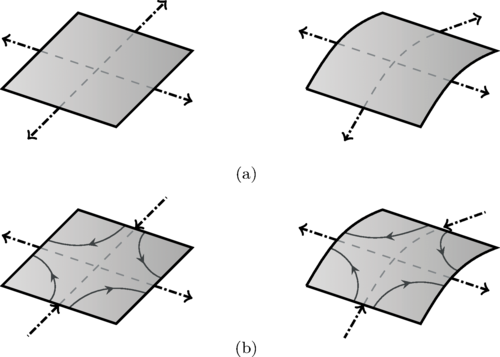
Lipid membranes are unusual materials. Lipids flow in-plane as a two-dimensional viscous fluid while the membrane bends out-of-plane as an elastic shell. The coupling between these two behaviors had been largely overlooked. Using differential geometry and linear irreversible thermodynamics, I developed a continuum theory for membrane dynamics across planar, spherical, and cylindrical geometries. A scaling analysis identified two governing dimensionless numbers: the Föppl-von Kármán number, comparing tension to bending forces, and the Scriven-Love number, a novel ratio comparing out-of-plane forces from intramembrane viscous stresses to elastic bending forces. Evaluating these numbers across physiological processes and in vitro experiments showed that in-plane viscosity cannot generally be neglected.
Key paper: Physical Review E 2020