Projects
Selected software projects. Full list on GitHub.

OpenADMET-ExpansionRx
Multi-architecture benchmarking pipeline for predicting drug absorption, distribution, metabolism, and toxicity (ADMET). Spans gradient-boosted trees, message-passing neural networks, and foundation models. Placed 25th of 100 finalists in the OpenADMET-ExpansionRx blind challenge across 9 endpoints. Uses clustering-based scaffold splits, temporal splits, and statistical hypothesis testing for architecture selection.
DDPM-Enhanced-Sampling
Generative framework using denoising diffusion probabilistic models (DDPMs) to sample polymer conformations consistent with Boltzmann statistics. Replaces expensive enhanced-sampling simulations with a learned model that generates physically valid conformations orders of magnitude faster.
Swimming-in-Potential-Flow
Companion code for the Journal of Fluid Mechanics paper on self-propulsion in potential flow. Implements boundary integral methods and analytical swimming theory in C++/CUDA for studying collective hydrodynamic interactions, demonstrating that viscous dissipation is not required for self-propulsion.
Analysis-Polyelectrolyte-Surface-Adsorption
High-performance analysis pipeline for polyelectrolyte molecular dynamics trajectories. Custom multi-threaded MDAnalysis extensions achieve 100x speedup over serial processing for enhanced-sampling simulations. Automates free-energy surface construction from biased trajectory data.

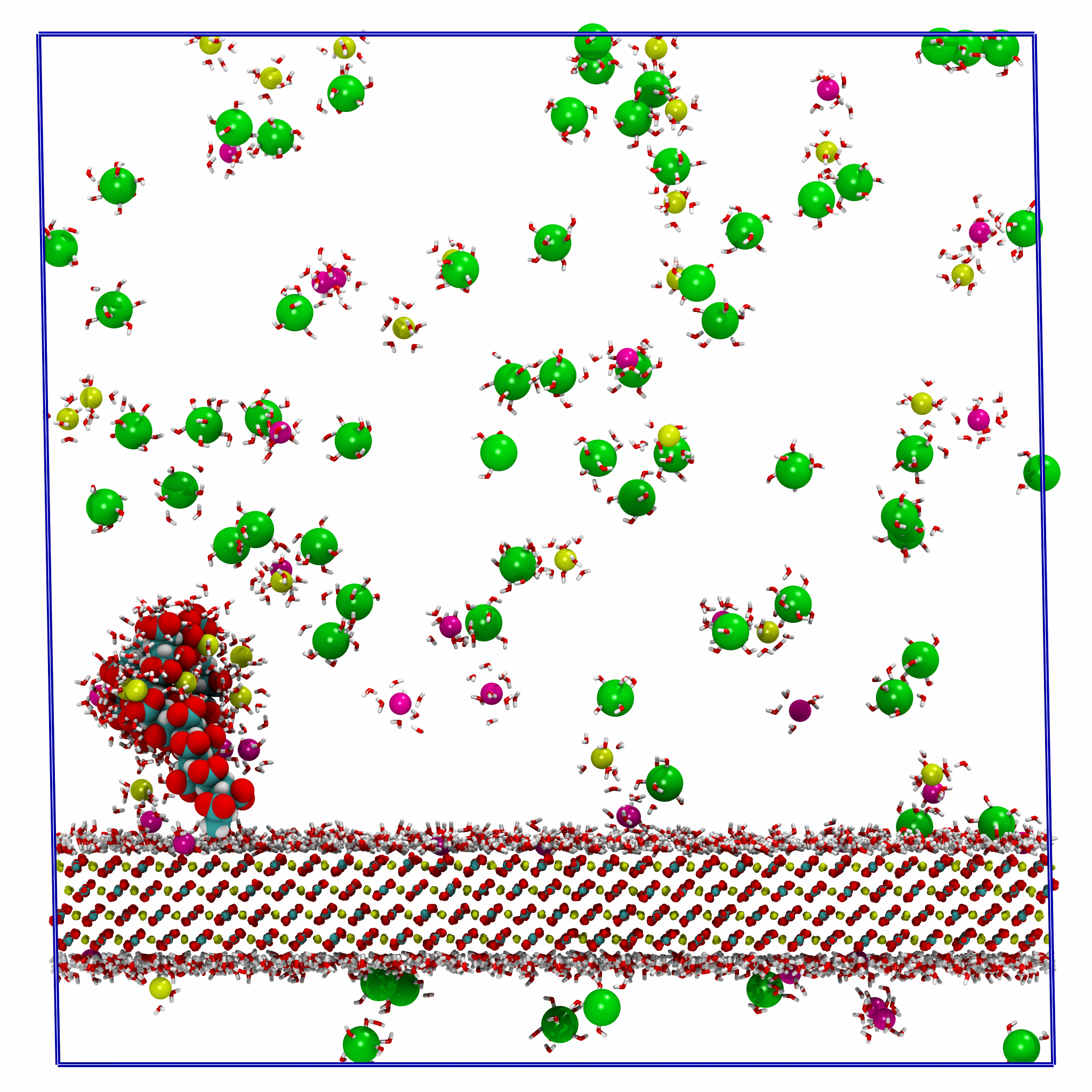
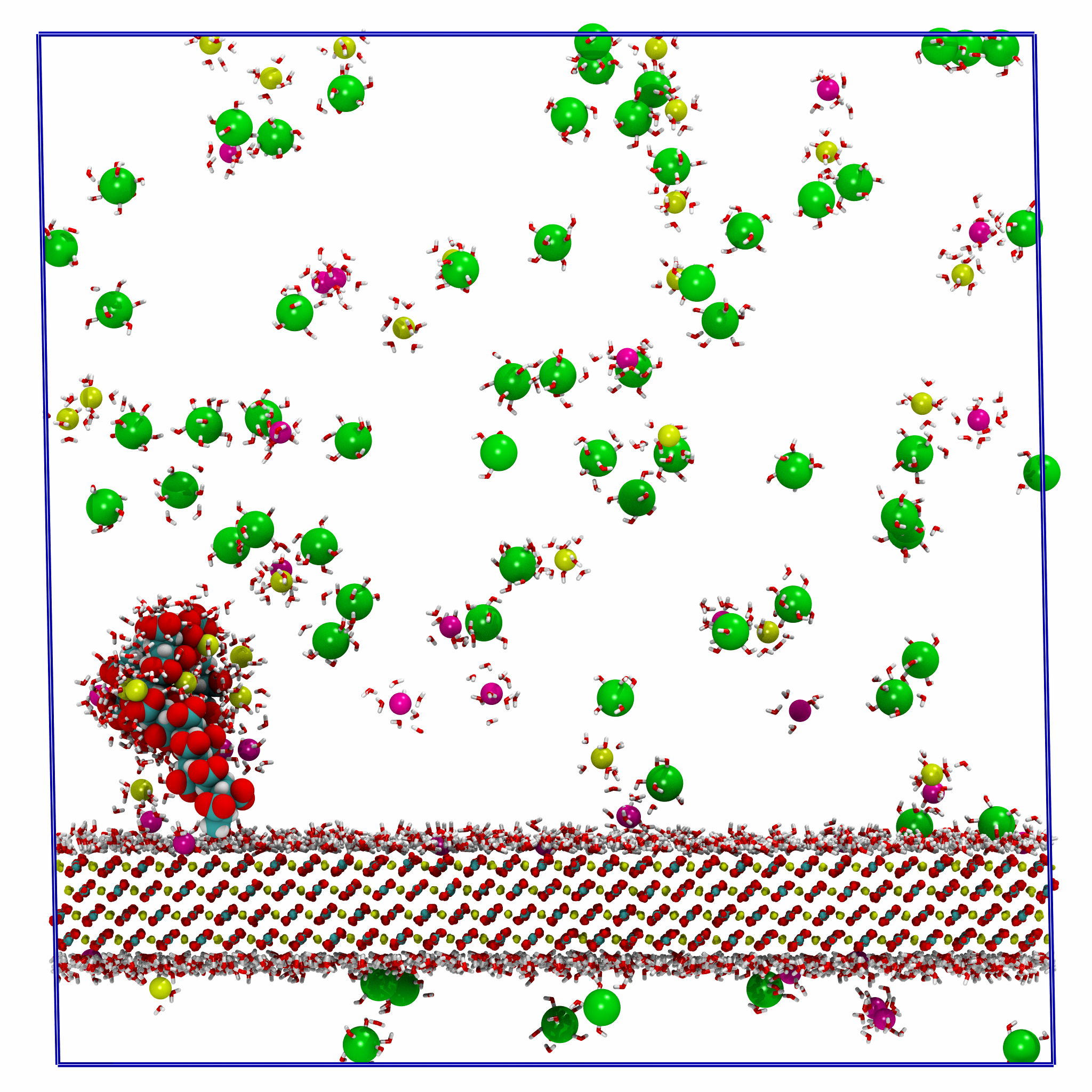
Polyelectrolyte-Surface-Adsorption
Production MD simulation workflows for studying polyelectrolyte adsorption to mineral surfaces. Uses GROMACS and metadynamics to compare direct and water-mediated binding modes on CaCO₃, with applications to water treatment and scale inhibition.
